Abstract
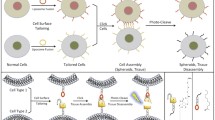
The ability to control cell patterning on artificial substrates with various physicochemical properties is of essence for important implications in cytology and biomedical fields. Despite extensive progress, the ability to control the cell-surface interaction is complicated by the complexity in the physiochemical features of bioactive surfaces. In particular, the manifestation of special wettability rendered by the combination of surface roughness and surface chemistry further enriches the cell-surface interaction. Herein we investigated the cell adhesion behaviors of Circulating Tumor Cells (CTCs) on topographically patterned but chemically homogeneous surfaces. Harnessing the distinctive cell adhesion on surfaces with different topography, we further explored the feasibility of controlled cell patterning using periodic lattices of alternative topographies. We envision that our method provides a designer’s toolbox to manage the extracellular environment.
Similar content being viewed by others
References
Barron J A, Wu P, Ladouceur H D, Ringeisen B R. Biological laser printing: A novel technique for creating heterogeneous 3-dimensional cell patterns. Biomedical Microdevices, 2004, 6, 139–147.
Díaz-Mochón J J, Tourniaire G, Bradley M. Microarray platforms for enzymatic and cell-based assays. Chemical Society Reviews, 2007, 36, 449–457.
Woodruff K, Fidalgo L M, Gobaa S, Lutolf M P, Maerkl S J. Live mammalian cell arrays. Nature Methods, 2013, 10, 550–552.
Kuss S, Polcari D, Geissler M, Brassard D, Mauzeroll J. Assessment of multidrug resistance on cell coculture patterns using scanning electrochemical microscopy. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110, 9249–9254.
Tan J L, Tien J, Pirone D M, Gray D S, Bhadriraju K, Chen C S. Cells lying on a bed of microneedles: An approach to isolate mechanical force. Proceedings of the National Academy of Sciences of the United States of America, 2003, 100, 1484–1489.
Qin Y P, Chen J, Bi Y, Xu X H, Zhou H, Gao J M, Hu Y, Zhao Y L, Chai Z F. Near-infrared light remote-controlled intracellular anti-cancer drug delivery using thermo/pH sensitive nanovehicle. Acta Biomaterialia, 2015, 17, 201–209.
Gao H, Bi Y, Chen J, Peng L R, Wen K K, Ji P, Ren W F, Li X Q, Zhang N, Gao J M, Chai Z F, Hu Y. Near-infrared light-triggered switchable nanoparticles for targeted chemo/photothermal cancer therapy. ACS Applied Materials & Interfaces, 2016, 8, 15103–15112.
Chen G C, Xie Y S, Peltier R, Lei H P, Wang P, Chen J, Hu Y, Wang F, Yao X, Sun H Y. Peptide-decorated gold nanoparticles as functional nano-capping agent of mesoporous silica container for targeting drug delivery. ACS Applied Materials & Interfaces, 2016, 8, 11204–11209.
Zhou H, Bi Y, Gao H, Chen P, Chen J, Hu Y. Reduction sensitive micellar magnetic nanoparticles for cancer targeted chemotherapy. Nanomedicine-Nanotechnology Biology and Medicine, 2016, 12, 547–547.
Qiao Y, An J, Ma L. Single cell array based assay for in vitro genotoxicity study of nanomaterials. Analytical Chemistry, 2013, 85, 4107–4112.
Heng L, Hu R, Chen S, Li M, Jiang L, Tang B Z. Ordered honeycomb structural interfaces for anticancer cells growth. Langmuir, 2013, 29, 14947–14953.
Giepmans B N G. The fluorescent toolbox for assessing protein location and function. Science, 2006, 312, 217–224.
Fernandes T G, Diogo M M, Clark D S, Dordick J S, Cabral J M S. High-throughput cellular microarray platforms: Applications in drug discovery, toxicology and stem cell research. Trends in Biotechnology, 2009, 27, 342–349.
Wang Y, Shah P, Phillips C, Sims C E, Allbritton N L. Trapping cells on a stretchable microwell array for single-cell analysis. Analytical and Bioanalytical Chemistry, 2011, 402, 1065–1072.
Liu X L, Chen L, Liu H L, Yang G, Zhang P C, Han D, Wang S T, Jiang L. Bio-inspired soft polystyrene nanotube substrate for rapid and highly efficient breast cancer-cell capture. NPG Asia Materials, 2013, 5, e63.
Zhang P, Chen L, Xu T, Liu H, Liu X, Meng J, Yang G, Jiang L, Wang S. Programmable fractal nanostructured interfaces for specific recognition and electrochemical release of cancer cells. Advanced Materials, 2013, 25, 3566–3570.
Liu H, Liu X, Meng J, Zhang P, Yang G, Su B, Sun K, Chen L, Han D, Wang S, Jiang L. Hydrophobic interactionmediated capture and release of cancer cells on thermoresponsive nanostructured surfaces. Advanced Materials, 2013, 25, 922–927.
Liu H L, Li Y Y, Sun K, Fan J B, Zhang P C, Meng J X, Wang S T, Jiang L. Dual-responsive surfaces modified with phenylboronic acid-containing polymer brush to reversibly capture and release cancer cells. Journal of the American Chemical Society, 2013, 135, 7603–7609.
Liu X, Wang S. Three-dimensional nano-biointerface as a new platform for guiding cell fate. Chemical Society Reviews, 2014, 43, 2385–2401.
Li Y Y, Lu Q H, Liu H L, Wang J F, Zhang P C, Liang H G, Jiang L, Wang S T. Antibody-modified reduced graphene oxide films with extreme sensitivity to circulating tumor cells. Advanced Materials, 2015, 27, 6848–6854.
Li G N, Yang G, Zhang P C, Li Y Y, Meng J X, Liu H L, Wang S T. Rapid cell patterning induced by differential topography on silica nanofractal substrates. Small, 2015, 11, 5642–5646.
Zhang F, Jiang Y, Liu X, Meng J, Zhang P, Liu H, Yang G, Li G, Jiang L, Wan L J, Hu J S, Wang S. Hierarchical nanowire arrays as three-dimensional fractal nanobiointerfaces for high efficient capture of cancer cells. Nano Letters, 2016, 16, 766–772.
Huang X W, Yue W Q, Liu D D, Yue J B, Li J Q, Sun D, Yang M, S Wang Z K. Monitoring the intracellular calcium response to a dynamic hypertonic environment. Scientific Reports, 2016, 6, 23591.
Deutsch M, Deutsch A, Shirihai O, Hurevich I, Afrimzon E, Shafran Y, Zurgil N. A novel miniature cell retainer for correlative high-content analysis of individual untethered non-adherent cells. Lab on a Chip, 2006, 6, 995–1000.
Wang X, Chen S, Kong M, Wang Z, Costa K D, Li R A, Sun D. Enhanced cell sorting and manipulation with combined optical tweezer and microfluidic chip technologies. Lab on a Chip, 2011, 11, 3656–3662.
Grier D G. A revolution in optical manipulation. Nature, 2003, 424, 810–816.
Liu W, Dechev N, Foulds I G, Burke R, Parameswaran A, Park E J. A novel permalloy based magnetic single cell micro array. Lab on a Chip, 2009, 9, 2381–2390.
Carlo D D, Wu L Y, Lee L P. Dynamic single cell culture array. Lab on a Chip, 2006, 6, 1445–1449.
Taff B M, Voldman J. A scalable addressable positivedielectrophoretic cell-sorting array. Analytical Chemistry, 2005, 77, 7976–7983.
Shi J, Ahmed D, Mao X, Lin S-C S, Lawit A, Huang T J. Acoustic tweezers: Patterning cells and microparticles using Standing Surface Acoustic Waves (SSAW). Lab on a Chip, 2009, 9, 2890–2895.
Rozkiewicz D I, Kraan Y, Werten M W T, de Wolf F A, Subramaniam V, Ravoo B J, Reinhoudt D N. Covalent microcontact printing of proteins for cell patterning. Chemistry-A European Journal, 2006, 12, 6290–6297.
Bearinger J, Dugan L, Wu L, Hill H, Christian A, Hubbell J. Chemical tethering of motile bacteria to silicon surfaces. BioTechniques, 2009, 46, 209–216.
Jang K, Sato K, Mawatari K, Konno T, Ishihara K, Kitamori T. Surface modification by 2-methacryloyloxyethyl phosphorylcholine coupled to a photolabile linker for cell micropatterning. Biomaterials, 2009, 30, 1413–1420.
Pan H-A, Hung Y-C, Su C-W, Tai S-M, Chen C-H, Ko F-H, Steve Huang G. A nanodot array modulates cell adhesion and induces an apoptosis-like abnormality in NIH-3Tcells. Nanoscale Research Letters, 2009, 4, 903–912.
Zhu H, Stybayeva G, Silangcruz J, Yan J, Ramanculov E, Dandekar S, George M D, Revzin A. Detecting cytokine release from single T-cells. Analytical Chemistry, 2009, 81, 8150–8156.
Wang Z, Chin S Y, Chin C D, Sarik J, Harper M, Justman J, Sia S K. Microfluidic CD4T-cell counting device using chemiluminescence-based detection. Analytical Chemistry, 2010, 82, 36–40.
Qiao Y, Wang C, Su M, Ma L. Single cell DNA damage/repair assay using HaloChip. Analytical Chemistry, 2012, 84, 1112–1116.
Hsiao S C, Liu H, Holstlaw T A, Liu C, Francis C Y, Francis M B. Real time assays for quantifying cytotoxicity with single cell resolution. PLOS ONE, 2013, 8, e66739.
Chen W, Weng S, Zhang F, Allen S, Li X, Bao L, Lam R H W, Macoska J A, Merajver S D, Fu J. Nanoroughened surfaces for efficient capture of circulating tumor cells without using capture antibodies. ACS Nano, 2013, 7, 566–575.
Dou X, Zhang D, Feng C, Jiang L. Bioinspired hierarchical surface structures with tunable wettability for regulating bacteria adhesion. ACS Nano, 2015, 9, 10664–10672.
Ranella A, Barberoglou M, Bakogianni S, Fotakis C, Stratakis E. Tuning cell adhesion by controlling the roughness and wettability of 3D micro/nano silicon structures. Acta Biomaterialia, 2010, 6, 2711–2720.
Yang G, Liu H, Liu X, Zhang P, Huang C, Xu T, Jiang L, Wang S. Underwater-transparent nanodendritic coatings for directly monitoring cancer cells. Advanced Healthcare Materials, 2013, 3, 332–337.
Liu H, Liu X, Meng J, Zhang P, Yang G, Su B, Sun K, Chen L, Han D, Wang S, Jiang L. Hydrophobic interactionmediated capture and release of cancer cells on thermoresponsive nanostructured surfaces. Advanced Materials, 2012, 25, 922–927.
Chen W, Sun Y, Fu J. Microfabricated nanotopological surfaces for study of adhesion-dependent cell mechanosensitivity. Small, 2012, 9, 81–89.
Kwak M, Han L, Chen J J, Fan R. Interfacing inorganic nanowire arrays and living cells for cellular function analysis. Small, 2015, 11, 5600–5610.
Feng W, Li L, Ueda E, Li J, Heißler S, Welle A, Trapp O, Levkin P A. Surface patterning via thiol-yne click chemistry: An extremely fast and versatile approach to superhydrophilic-superhydrophobic micropatterns. Advanced Materials Interfaces, 2014, 1, 1400269.
Wang L, Asghar W, Demirci U, Wan Y. Nanostructured substrates for isolation of circulating tumor cells. Nano Today, 2013, 8, 374–387.
Chung S H, Son S J, Min J. The nanostructure effect on the adhesion and growth rates of epithelial cells with well-defined nanoporous alumina substrates. Nanotechnology, 2010, 21, 125104.
Lopacinska J M, Gradinaru C, Wierzbicki R, Kobler C, Schmidt M S, Madsen M T, Skolimowski M, Dufva M, Flyvbjerg H, Molhave K. Cell motility, morphology, viability and proliferation in response to nanotopography on silicon black. Nanoscale, 2012, 4, 3739–3745.
Zhang P, Lin L, Zang D, Guo X, Liu M. Designing bioinspired anti-biofouling surfaces based on a superwettability strategy. Small, 2017, 13, 1503334.
Kim D-J, Lee G, Kim G-S, Lee S-K. Statistical analysis of immuno-functionalized tumor-cell behaviors on nanopatterned substrates. Nanoscale Research Letters, 2012, 7, 637.
Geyer F L, Ueda E, Liebel U, Grau N, Levkin P A. Superhydrophobic-superhydrophilic micropatterning: Towards genome-on-a-chip cell microarrays. Angewandte Chemie International Edition, 2011, 50, 8424–8427.
Cristofanilli M, Hayes D F, Budd G T, Ellis M J, Stopeck A, Reuben J M, Doyle G V, Matera J, Allard W J, Miller M C, Fritsche H A, Hortobagyi G N, Terstappen L. Circulating tumor cells: A novel prognostic factor for newly diagnosed metastatic breast cancer. Journal of Clinical Oncology, 2005, 23, 1420–1430.
Bailey S N, Ali S M, Carpenter A E, Higgins C O, Sabatini D M. Microarrays of lentiviruses for gene function screens in immortalized and primary cells. Nature Methods, 2006, 3, 117–122.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhou, X., Li, J., Sun, H. et al. Controlled cell patterning on bioactive surfaces with special wettability. J Bionic Eng 14, 440–447 (2017). https://doi.org/10.1016/S1672-6529(16)60409-2
Published:
Issue Date:
DOI: https://doi.org/10.1016/S1672-6529(16)60409-2